VueGen is a Python package that automates the creation of scientific reports.
| Information | Links |
|---|---|
| Package |     |
| Documentation |   |
| Build | |
| Examples |   |
| Discuss on GitHub |   |
| Cite |  |
- About the project
- Installation
- Execution
- Case studies
- Web application deployment
- Credits and acknowledgements
- Citation
- Contact and feedback
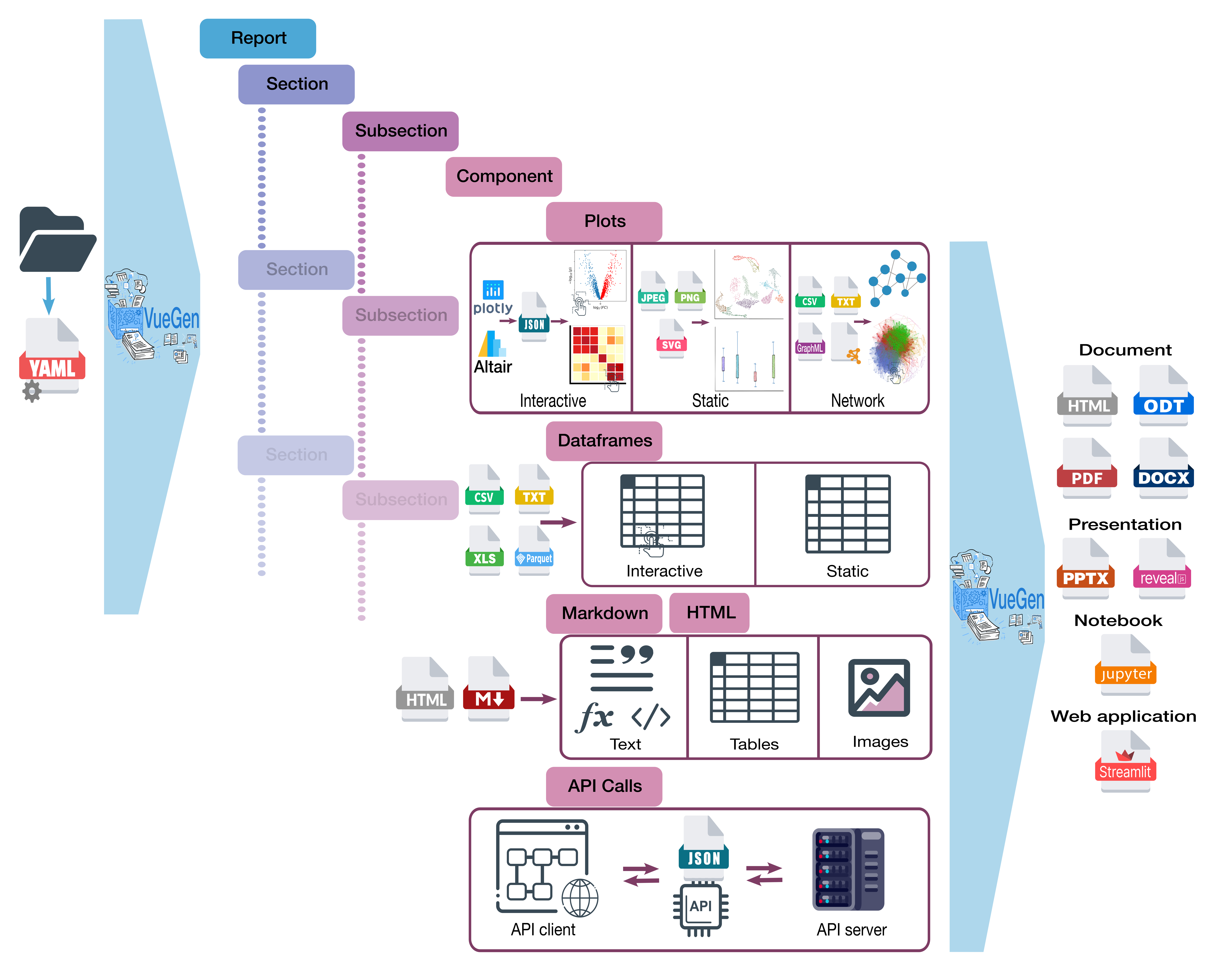
VueGen automates the creation of reports based on a directory with plots, dataframes, and other files in different formats. A YAML configuration file is generated from the directory to define the structure of the report. Users can customize the report by modifying the configuration file, or they can create their own configuration file instead of passing a directory as input.
The configuration file specifies the structure of the report, including sections, subsections, and various components such as plots, dataframes, markdown, html, and API calls. Reports can be generated in various formats, including documents (PDF, HTML, DOCX, ODT), presentations (PPTX, Reveal.js), notebooks (Jupyter) or Streamlit web applications.
An overview of the VueGen workflow is shown in the figure below:
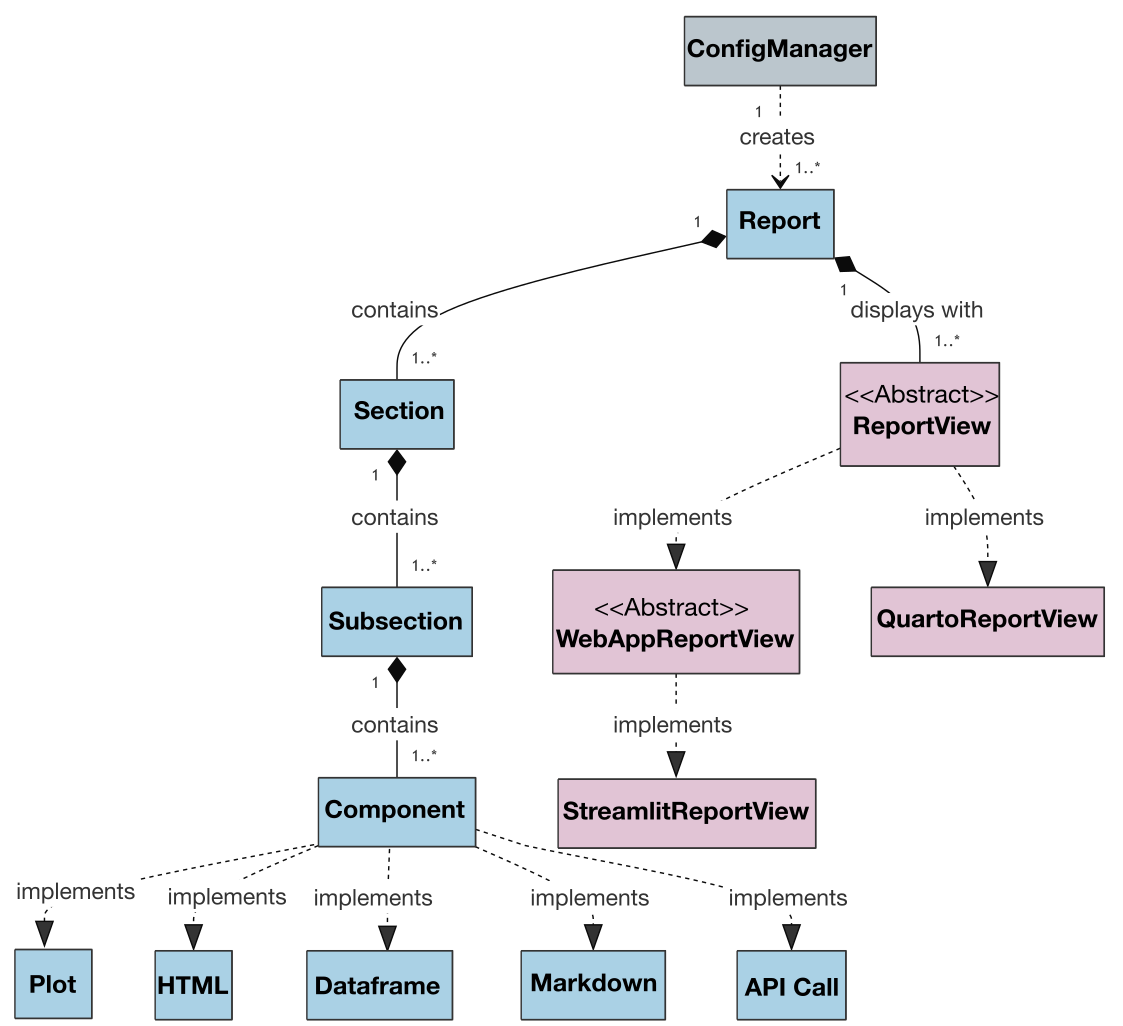
Also, the class diagram for the project is presented below to illustrate the architecture and relationships between classes:
An extended version of the class diagram with attributes and methods is available here.
The VueGen documentation is available at vuegen.readthedocs.io, where you can find detailed information of the package’s classes and functions, installation and execution instructions, and case studies to demonstrate its functionality.
VueGen is available on PyPI and can be installed using pip:
pip install vuegenYou can also install the package for development by cloning this repository and running the following command:
pip install -e path/to/vuegen # specify location
pip install -e . # in case your pwd is in the vuegen directoryTip
It is recommended to install VueGen inside a virtual environment to manage depenendencies and avoid conflicts with existing packages. You can use the virtual environment manager of your choice, such as poetry, conda, or pipenv.
VueGen is also available on Bioconda and can be installed using conda:
conda install bioconda::vuegenVueGen uses Quarto to generate various report types. The pip insallation includes quarto using the quarto-cli Python library. To test if quarto is installed in your computer, run the following command:
quarto checkTip
If quarto is not installed, you can download the command-line interface from the Quarto website for your operating system.
If you prefer not to install VueGen on your system, a pre-configured Docker container is available. It includes all dependencies, ensuring a fully reproducible execution environment. See the Execution section for details on running VueGen with Docker. The official Docker image is available at quay.io/dtu_biosustain_dsp/vuegen.
VueGen is also available as a nf-core module, customised for compatibility with the Nextflow environment. This module is designed to automate report generation from outputs produced by other modules, subworkflows, or pipelines. The code and documentation for the nf-core module are available in the nf-VueGen repository.
Important
Here we use the Earth_microbiome_vuegen_demo_notebook directory and the Earth_microbiome_vuegen_demo_notebook.yaml configuration file as examples, which are available in the docs/example_data and docs/example_config_files folders, respectively. Make sure to clone this reposiotry to access these contents, or use your own directory and configuration file.
Run VueGen using a directory with the following command:
vuegen --directory docs/example_data/Earth_microbiome_vuegen_demo_notebook --report_type streamlitNote
By default, the streamlit_autorun argument is set to False, but you can use it in case you want to automatically run the streamlit app.
It's also possible to provide a configuration file instead of a directory:
vuegen --config docs/example_config_files/Earth_microbiome_vuegen_demo_notebook.yaml --report_type streamlitThe current report types supported by VueGen are:
- Streamlit
- HTML
- DOCX
- ODT
- Reveal.js
- PPTX
- Jupyter
Instead of installing VueGen locally, you can run it directly from a Docker container with the following command:
docker run --rm \
-v "$(pwd)/docs/example_data/Earth_microbiome_vuegen_demo_notebook:/home/appuser/Earth_microbiome_vuegen_demo_notebook" \
-v "$(pwd)/output_docker:/home/appuser/quarto_report" \
quay.io/dtu_biosustain_dsp/vuegen:docker --directory /home/appuser/Earth_microbiome_vuegen_demo_notebook --report_type streamlitWe have a simple GUI for VueGen that can be run locally or through a standalone executable.
cd gui
python app.pyThe bundle GUI with the VueGen package is available under the releases. You will need to
unzip the file and run vuegen_gui in the unpacked main folder. Most dependencies are included into
the bundle under _internals using PyInstaller.
Streamlit works out of the box as a purely Python based package. For html creation you will have to
have a global Python installation with the jupyter package installed. quarto needs to start
a kernel for execution. This is also true if you install quarto globally on your machine.
More information can be found in the GUI README.
VueGen’s functionality is demonstrated through two case studies:
1. Predefined Directory
This introductory case study uses a predefined directory with plots, dataframes, Markdown, and HTML components. Users can generate reports in different formats and modify the configuration file to customize the report structure.
2. Earth Microbiome Project Data
This advanced case study demonstrates the application of VueGen in a real-world scenario using data from the Earth Microbiome Project (EMP). The EMP is an initiative to characterize global microbial taxonomic and functional diversity. The notebook process the EMP data, create plots, dataframes, and other components, and organize outputs within a directory to produce reports. Report content and structure can be adapted by modifying the configuration file. Each report consists of sections on exploratory data analysis, metagenomics, and network analysis.
Once a Streamlit report is generated, it can be deployed as a web application to make it accessible online. There are multiple ways to achieve this:
- Streamlit Community Cloud: Deploy your report easily using Streamlit Cloud, as demonstrated in the EMP VueGen Demo. The process involves moving the necessary scripts, data, and a requirements.txt file into a GitHub repository. Then, the app can be deployed via the Streamlit Cloud interface. The deployment example is available in the
streamlit-report-examplebranch. - Standalone Executables: Convert your Streamlit application into a desktop app by packaging it as an executable file for different operating systems. A detailed explanation of this process can be found in this Streamlit forum post.
- Stlite: Run Streamlit apps directly in the browser with stlite, a WebAssembly port of Streamlit powered by Pyodide, eliminating the need for a server. It also allows packaging apps as standalone desktop executables using stlite desktop.
These options provide flexibility depending on whether the goal is online accessibility, lightweight execution, or local application distribution.
- VueGen was developed by the Multiomics Network Analytics Group (MoNA) at the Novo Nordisk Foundation Center for Biosustainability (DTU Biosustain).
- VueGen relies on the work of numerous open-source projects like Streamlit, Quarto, and others. A big thank you to their authors for making this possible!
- The vuegen logo was designed based on an image created by Scriberia for The Turing Way Community, which is shared under a CC-BY licence. The original image can be found at Zenodo.
If you use VueGen in your research or publications, please cite it as follows:
APA:
Ayala-Ruano, S., Webel, H., & Santos, A. (2025). VueGen: Automating the generation of scientific reports. bioRxiv. https://doi.org/10.1101/2025.03.05.641152
BibTeX:
@article{Ayala-Ruano2025VueGen,
author = {Ayala-Ruano, Sebastian and Webel, Henry and Santos, Alberto},
title = {VueGen: Automating the generation of scientific reports},
journal = {bioRxiv},
year = {2025},
doi = {10.1101/2025.03.05.641152},
publisher = {Cold Spring Harbor Laboratory},
url = {https://www.biorxiv.org/content/10.1101/2025.03.05.641152},
eprint = {https://www.biorxiv.org/content/10.1101/2025.03.05.641152.full.pdf}
}We appreciate your feedback! If you have any comments, suggestions, or run into issues while using VueGen, feel free to open an issue in this repository. Your input helps us make VueGen better for everyone.