|
2 | 2 |
|
3 | 3 | [](https://badge.fury.io/py/tangram-sc) |
4 | 4 |
|
5 | | -Tangram is a Python package, written in [PyTorch](https://pytorch.org/) and based on [scanpy](https://scanpy.readthedocs.io/en/stable/), for mapping single-cell (or single-nucleus) gene expression data onto spatial gene expression data. The single-cell dataset and the spatial dataset should be collected from the same anatomical region/tissue type, ideally from a biological replicate, and need to share a set of genes. Tangram aligns the single-cell data in space by fitting gene expression on the shared genes. The best way to familiarize yourself with Tangram is to check out [our tutorial](https://github.com/broadinstitute/Tangram/blob/master/example/1_tutorial_tangram.ipynb). [](https://colab.research.google.com/drive/1SVLUIZR6Da6VUyvX_2RkgVxbPn8f62ge?usp=sharing) |
| 5 | +Tangram is a Python package, written in [PyTorch](https://pytorch.org/) and based on [scanpy](https://scanpy.readthedocs.io/en/stable/), for mapping single-cell (or single-nucleus) gene expression data onto spatial gene expression data. The single-cell dataset and the spatial dataset should be collected from the same anatomical region/tissue type, ideally from a biological replicate, and need to share a set of genes. Tangram aligns the single-cell data in space by fitting gene expression on the shared genes. The best way to familiarize yourself with Tangram is to check out [our tutorial](https://github.com/broadinstitute/Tangram/blob/master/tangram_tutorial.ipynb). [](https://colab.research.google.com/drive/1SVLUIZR6Da6VUyvX_2RkgVxbPn8f62ge?usp=sharing) |
6 | 6 |
|
7 | 7 | 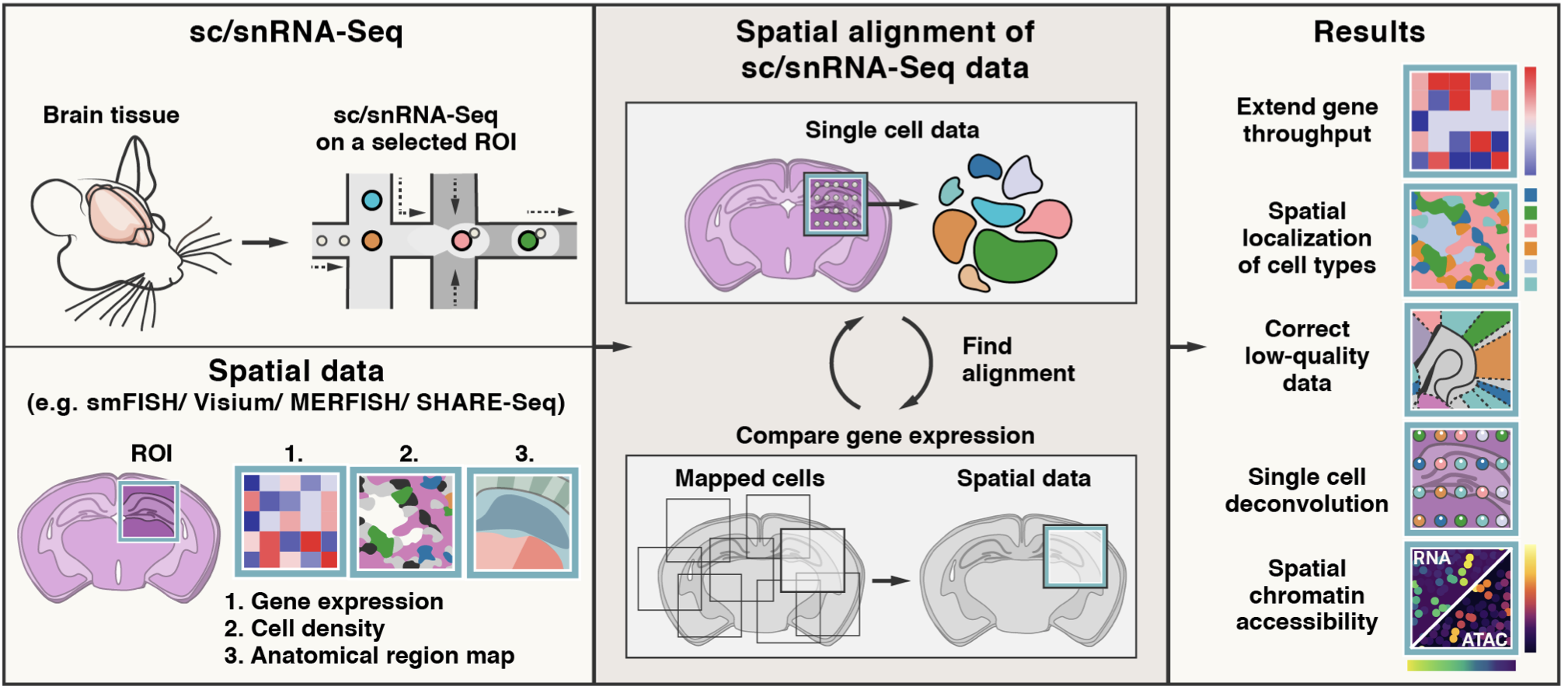 |
8 | 8 | Tangram has been tested on various types of transcriptomic data (10Xv3, Smart-seq2 and SHARE-seq for single cell data; MERFISH, Visium, Slide-seq, smFISH and STARmap as spatial data). In our [preprint](https://www.biorxiv.org/content/10.1101/2020.08.29.272831v1), we used Tangram to reveal spatial maps of cell types and gene expression at single cell resolution in the adult mouse brain. More recently, we have applied our method to different tissue types including human lung, human kidney developmental mouse brain and metastatic breast cancer. |
@@ -52,7 +52,7 @@ The returned AnnData,`ad_map`, is a cell-by-voxel structure where `ad_map.X[i, j |
52 | 52 |
|
53 | 53 | The returned `ad_ge` is a voxel-by-gene AnnData, similar to spatial data `ad_sp`, but where gene expression has been projected from the single cells. This allows to extend gene throughput, or correct for dropouts, if the single cells have higher quality (or more genes) than single cell data. It can also be used to transfer cell types onto space. |
54 | 54 |
|
55 | | -For more details on how to use Tangram check out [our tutorial](https://github.com/broadinstitute/Tangram/blob/master/example/1_tutorial_tangram.ipynb). [](https://colab.research.google.com/drive/1SVLUIZR6Da6VUyvX_2RkgVxbPn8f62ge?usp=sharing) |
| 55 | +For more details on how to use Tangram check out [our tutorial](https://github.com/broadinstitute/Tangram/blob/master/tangram_tutorial.ipynb). [](https://colab.research.google.com/drive/1SVLUIZR6Da6VUyvX_2RkgVxbPn8f62ge?usp=sharing) |
56 | 56 |
|
57 | 57 | *** |
58 | 58 |
|
@@ -126,4 +126,4 @@ PyPI maintainer: |
126 | 126 | |
127 | 127 |
|
128 | 128 | The artwork has been curated by: |
129 | | -- Anna Hupalowska <[email protected]> |
| 129 | +- Anna Hupalowska <[email protected]> |
0 commit comments