IBBI is a Python package that provides a simple and unified interface for detecting and classifying bark and ambrosia beetles from images using state-of-the-art computer vision models.
This package is designed to support entomological research by automating the laborious task of beetle identification, enabling high-throughput data analysis for ecological studies, pest management, and biodiversity monitoring.
The ability to accurately detect and identify bark and ambrosia beetles is critical for forest health and pest management. However, traditional methods face significant challenges:
- They are slow and time-consuming.
- They require highly specialized expertise.
- They create a bottleneck for large-scale research.
The IBBI package provides a powerful, modern solution to overcome these obstacles by making available pre-trained, open-source models to automate detection and classification from images, lowering the barrier to entry for researchers. The IBBI package also makes other advanced techniques available to the user such as general zero-shot detection, general feature extraction, model evaluation, and model explainability.
-
Model Access:
Access powerful models with a single function callibbi.create_model(). The following types of models are available:-
Single-Class Bark Beetle Detection: Detect the presence of any bark beetle in an image. These models have been trained with the task of single-class object detection on Bark and ambrosia beetle specific data. These models do not classify the species of beetle, only its presence and location in an image.
-
Multi-Class Species Detection: Identify the species of a beetle from an image. These models have been trained with the task of multi-class object detection on Bark and ambrosia beetle specific data. These models classify the species of beetle, as well as its presence and location in an image.
-
General Zero-Shot Detection: Detect objects using a text prompt (e.g., "insect"), without prior training on bark and ambrosia beetle specific data (e.g., GroundingDINO, YoloWorld). These models have been trained on large, diverse datasets and can generalize to detect a wide range of objects based on textual descriptions.
-
General Pre-trained Feature Extraction: Extract feature embeddings from images using pre-trained models without additional training for specifically identifying bark and ambrosia beetles (e.g., DINOv3, EVA-02).
-
-
Model Evaluation:
This package includes a small set of ~2 000 images of 63 species for benchmarking model performance using a simple call to theibbi.Evaluator()wrapper. The evaluation wrapper makes computing of the following types of metrics easy:-
Classification Metrics: Assesses the model's ability to correctly classify species. The
evaluator.classification()method returns a comprehensive set of metrics including accuracy, balanced accuracy, F1-score, Cohen's Kappa, precision, and recall alongside a confusion matrix. -
Object Detection Metrics: Evaluates the accuracy of bounding box predictions. The
evaluator.object_detection()method calculates the mean Average Precision (mAP) over a range of Intersection over Union (IoU) thresholds. You can also get per-class AP scores to understand performance on a species-by-species basis. -
Embedding & Clustering Metrics: Evaluates the quality of the feature embeddings generated by the models. The
evaluator.embeddings()method performs dimensionality reduction with Uniform Manifold Approximation and Projection (UMAP) and clustering with Hierarchical Density-Based Spatial Clustering of Applications with Noise (HDBSCAN), then calculates:- Intrinsic metrics to assess the quality of the clusters themselves, such as the Silhouette Score, Davies-Bouldin Index, and Calinski-Harabasz Index.
- Extrinsic metrics that compare the clusters to the ground-truth species labels, including the Adjusted Rand Index (ARI), Normalized Mutual Information (NMI), and Cluster Purity.
- Mantel Correlation: A Mantel test is performed to see if the distances between species in the embedding space correlate with a known external distance matrix (e.g., taxonomic/phylogenetic distance). By default the package uses a sample phylogenetic distance matrix constrained to the current taxonomy of the species with branch lengths based on evolutionary divergence times of the COI gene to act as an estimate of evolutionary divergence.
-
-
Model Explainability:
Gain insights into why a model makes certain predictions with integrated explainability methods. Theibbi.Explainer()wrapper provides a simple interface for two popular techniques:- SHapley Additive exPlanations (SHAP): A powerful, game theory-based approach that attributes a prediction to the features of an input. The
explainer.with_shap()method generates robust, theoretically-grounded explanations for a set of images, which is ideal for understanding the contribution of each part of an image to a prediction. - Local Interpretable Model-agnostic Explanations (LIME): A technique that explains the predictions of any classifier by learning an interpretable model locally around the prediction. Use the
explainer.with_lime()method for a quicker, more intuitive visualization of which parts of a single image were most influential in the model's decision.
- SHapley Additive exPlanations (SHAP): A powerful, game theory-based approach that attributes a prediction to the features of an input. The
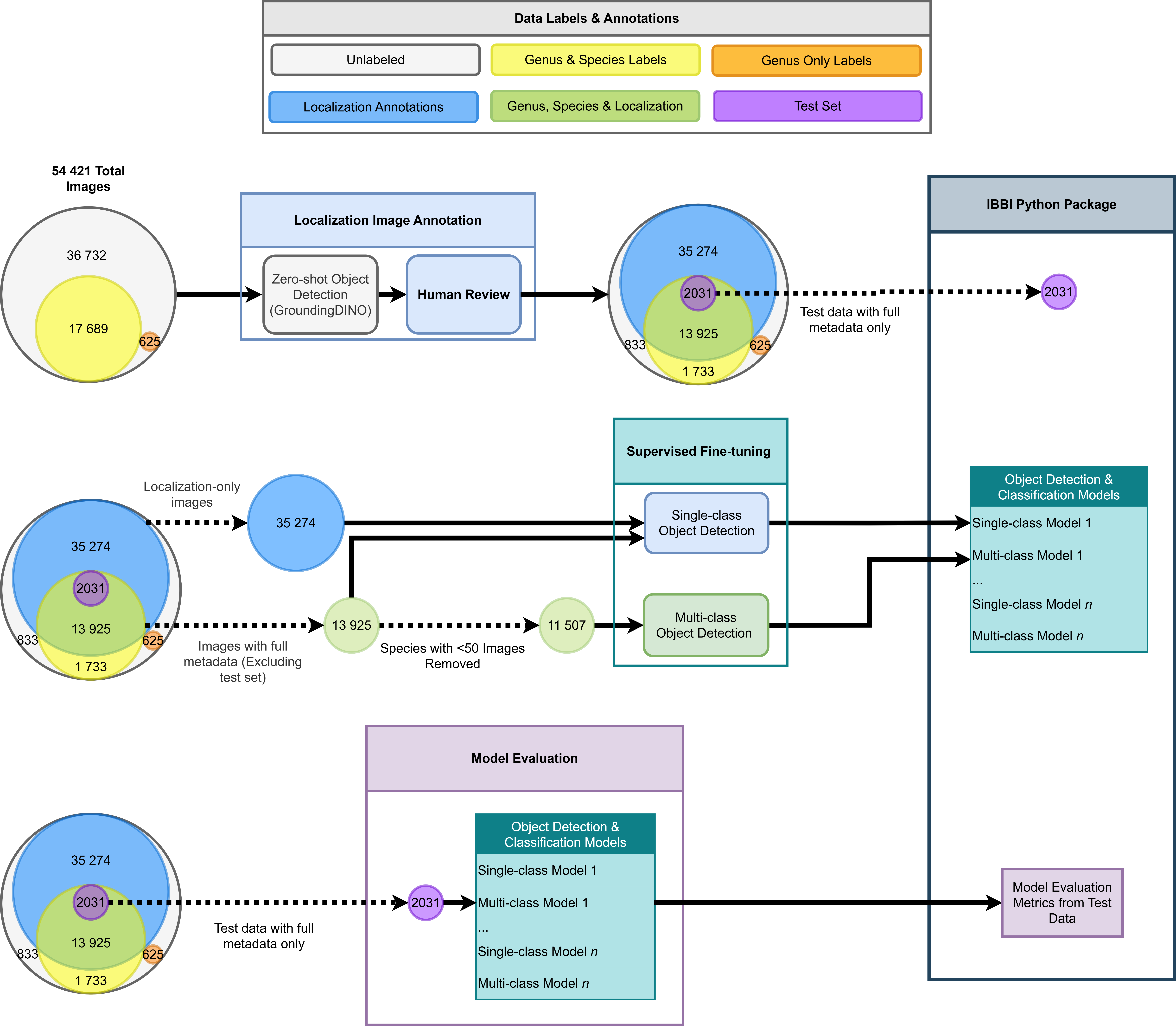
The trained models in ibbi are the result of a comprehensive data collection, annotation, and training pipeline by the Forest Entomology Lab at the University of Florida.
-
Data Collection and Curation: The process begins with data collection from various sources. A zero-shot detection model performs initial bark beetle localization, followed by human-in-the-loop verification to ensure accurate bounding box annotations. Species classification is performed by expert taxonomists to provide high-quality species labels.
-
Model-Specific Training Data: The annotated dataset is curated for different model types:
- Single-Class Detection: Trained in a supervised manner on the task of object-detection and object-classification using all images with verified bark beetle localizations.
- Multi-Class Species Detection: Trained in a supervised manner on the task of object-detection and object-classification using images with both verified localizations and species-level labels. To ensure robustness, species with fewer than 50 images are excluded.
Note - No additional training is performed for the zero-shot detection and general feature extraction models, as they leverage pre-trained weights from large, diverse datasets.
-
Evaluation and Deployment: A held-out test set is used to evaluate all models. A summary of performance metrics can be viewed with
ibbi.list_models()or by simply viewing themodel summary table. Alternatively evalutation can be done independently by runningibbi.Evaluator(). The trained models are stored under theIBBI-bio Hugging Face Hub communityfor easy access.
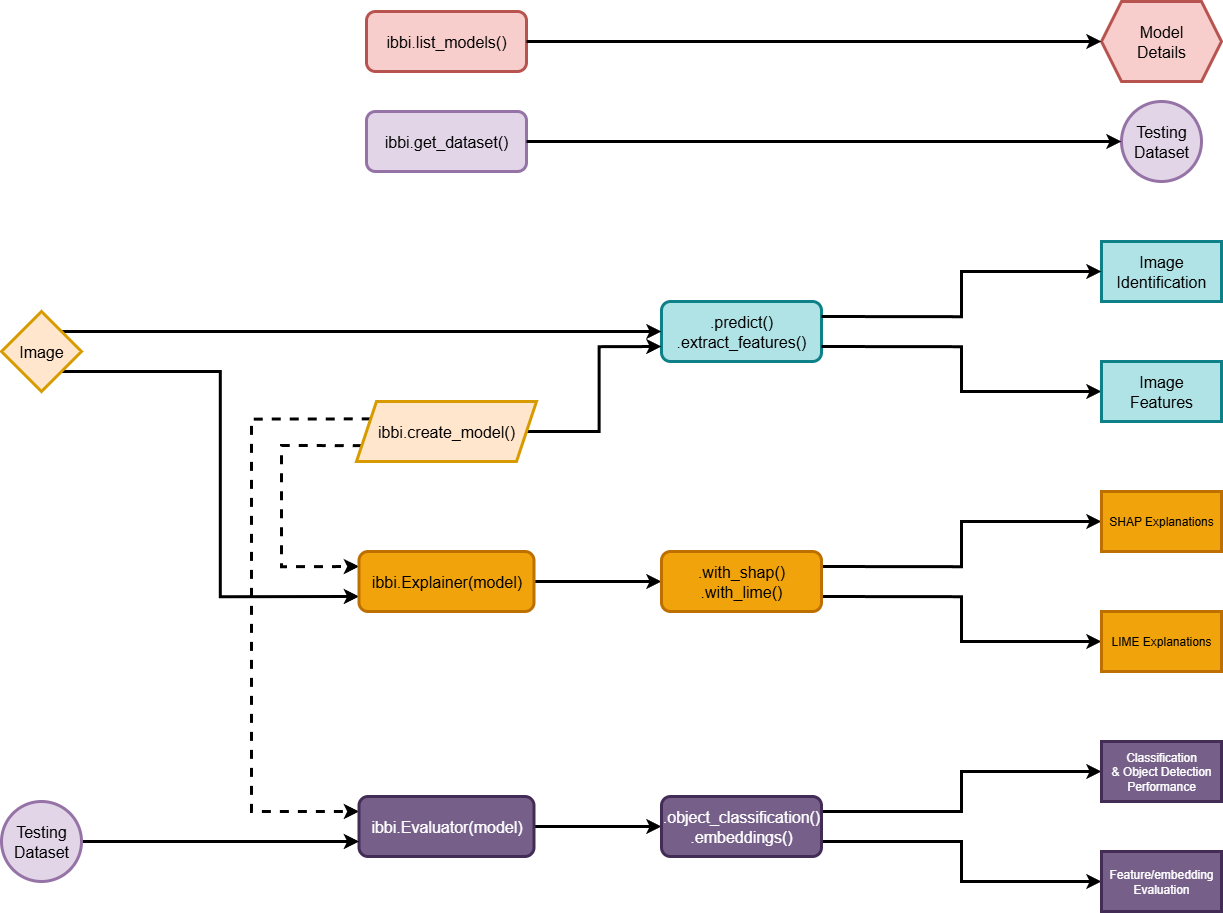
The ibbi package is designed to be simple and intuitive. The following diagram summarizes the main functions, classes and methods including their inputs and outputs.
This package requires PyTorch. For compatibility with your specific hardware (e.g., CUDA-enabled GPU), please install PyTorch before installing ibbi.
1. Install PyTorch
Follow the official instructions at pytorch.org to install the correct version for your system.
2. Install IBBI
Once PyTorch is installed, install the package from PyPI:
pip install ibbiOr install the latest development version directly from GitHub:
pip install git+https://github.com/ChristopherMarais/IBBI.git-
Disk Space: A minimum of 10-20 GB of disk space is recommended to accommodate the Python environment, downloaded models, and cached datasets.
-
CPU & RAM: Running inference on a CPU is possible but can be slow. For model evaluation on large datasets (like the built-in test set), a significant amount of RAM (16GB, 32GB, or more) is highly recommended to avoid memory crashes.
-
GPU (Recommended): A CUDA-enabled GPU (e.g., NVIDIA T4, RTX 3060 or better) with at least 8GB of VRAM is strongly recommended for both inference and model evaluation.
Using ibbi is straightforward. Load a model and immediately use it for inference.
import ibbi
# --- List Available Models ---
ibbi.list_models()
# --- Create a Model ---
classifier = ibbi.create_model(model_name="species_classifier", pretrained=True)
# --- Perform Inference ---
results = classifier.predict("path/to/your/image.jpg")For more detailed demonstrations, please see the example notebooks located in the notebooks/ folder of the repository.
To see a list of available models and their performance metrics directly from Python, run:
ibbi.list_models()Replace model_name with one of the available model names to use those models.
Model Summary Table
The most detailed version of the table can also be found here.
ID = In-Distribution (i.e., species seen during training) OOD = Out-of-Distribution (i.e., species not seen during training)
| Model Name | Tasks | Pretrained Weights Repository | Paper | Model Size (Parameters) | Image size | Embedding vector shape | # of images fine-tuned | Epochs fine-tuned | Species mAP (ID) | Genus mAP (ID) | Genus mAP (OOD) | Organism mAP (ID) | Organism mAP (OOD) | Silhouette (ID) | Davies-Bouldin (ID) | Calinski-Harabasz (ID) | ARI (ID) | NMI (ID) | Cluster Purity (ID) | Mantel R (ID) | p-value (ID) | Mantels R support (ID) | Silhouette (OOD) | Davies-Bouldin (OOD) | Calinski-Harabasz (OOD) | ARI (OOD) | NMI (OOD) | Cluster Purity (OOD) | Mantel R (OOD) | p-value (OOD) | Mantels R support (OOD) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| yoloworldv2_bb_detect_model | Zero-shot Object Detection (Prompt: 'insect'), Feature extraction | https://github.com/ultralytics/assets/releases/download/v8.3.0/yolov8x-worldv2.pt | https://arxiv.org/pdf/2401.17270v2 | 68.2M | 640x640 | ( , 640) | 0 | N/A | N/A | N/A | N/A | 0.1997 | 0.1684 | 0.3334 | 1.146 | 328.7954 | 0.0011 | 0.0212 | 0.0823 | -0.1506 | 0.311 | 32 | 0.2562 | 1.1183 | 99.6116 | 0.0024 | 0.0328 | 0.1115 | -0.071 | 0.482 | 72 |
| yolov9e_bb_multi_class_detect_model | Multi-class Object Detection, Feature extraction | https://huggingface.co/IBBI-bio/ibbi_yolov9_oc | https://arxiv.org/abs/2402.13616 | 58.1M | 640x640 | ( , 512) | 11 507 | 100 | 0.89 | 0.9371 | 0.1853 | 0.9745 | 0.7769 | 0.2797 | 0.8943 | 583.5217 | 0.0072 | 0.1272 | 0.1363 | -0.0395 | 0.749 | 32 | 0.2083 | 1.0573 | 39.5503 | 0.0029 | 0.0299 | 0.1757 | -0.0154 | 0.867 | 72 |
| yolov9e_bb_detect_model | Single-class Object Detection, Feature extraction | https://huggingface.co/IBBI-bio/ibbi_yolov9_od | https://arxiv.org/abs/2402.13616 | 58.1M | 640x640 | ( , 512) | 46 781 | 100 | N/A | N/A | N/A | 0.9609 | 0.9447 | 0.1714 | 1.4861 | 243.5296 | 0.0013 | 0.0302 | 0.0794 | -0.0637 | 0.657 | 32 | 0.2428 | 1.4458 | 741.849 | 0.0214 | 0.138 | 0.1956 | 0.0005 | 0.997 | 72 |
| yolov8x_bb_multi_class_detect_model | Multi-class Object Detection, Feature extraction | https://huggingface.co/IBBI-bio/ibbi_yolov8_oc | https://arxiv.org/abs/2408.15857 | 68.2M | 640x640 | ( , 640) | 11 507 | 100 | 0.8842 | 0.9255 | 0.2282 | 0.9731 | 0.7196 | 0.262 | 0.9425 | 587.5894 | 0.0046 | 0.1004 | 0.1156 | -0.1092 | 0.393 | 32 | 0.3593 | 1.1656 | 210.2135 | 0.0019 | 0.0251 | 0.1117 | -0.0886 | 0.202 | 72 |
| yolov8x_bb_detect_model | Single-class Object Detection, Feature extraction | https://huggingface.co/IBBI-bio/ibbi_yolov8_od | https://arxiv.org/abs/2408.15857 | 68.2M | 640x640 | ( , 640) | 46 781 | 100 | N/A | N/A | N/A | 0.9673 | 0.9559 | 0.0561 | 1.3672 | 163.8281 | 0.0043 | 0.0669 | 0.1013 | -0.0536 | 0.718 | 32 | 0.3316 | 1.2511 | 3302.8547 | -0.002 | 0.0187 | 0.1307 | 0.0094 | 0.922 | 72 |
| yolov12x_bb_multi_class_detect_model | Multi-class Object Detection, Feature extraction | https://huggingface.co/IBBI-bio/ibbi_yolov12_oc | https://arxiv.org/pdf/2502.12524 | 59.1M | 640x640 | ( , 768) | 12 507 | 100 | 0.8854 | 0.9371 | 0.2059 | 0.9691 | 0.7021 | 0.2657 | 1.0177 | 434.3505 | 0.0044 | 0.0937 | 0.1133 | -0.1194 | 0.382 | 32 | 0.2578 | 1.2137 | 45.696 | 0.0006 | 0.0097 | 0.1123 | -0.0419 | 0.641 | 72 |
| yolov12x_bb_detect_model | Single-class Object Detection, Feature extraction | https://huggingface.co/IBBI-bio/ibbi_yolov12_od | https://arxiv.org/pdf/2502.12524 | 59.1M | 640x640 | ( , 768) | 46 781 | 100 | N/A | N/A | N/A | 0.9716 | 0.7546 | 0.3258 | 1.1295 | 627.9548 | 0.0042 | 0.0852 | 0.1002 | -0.1532 | 0.279 | 32 | 0.2189 | 1.2177 | 28.533 | 0.0004 | 0.0069 | 0.1121 | 0.0643 | 0.466 | 72 |
| yolov11x_bb_multi_class_detect_model | Multi-class Object Detection, Feature extraction | https://huggingface.co/IBBI-bio/ibbi_yolov11_oc | https://www.arxiv.org/abs/2410.17725 | 56.9M | 640x640 | ( , 768) | 11 507 | 100 | 0.8641 | 0.9171 | 0.2006 | 0.9511 | 0.6939 | 0.287 | 1.1504 | 509.429 | 0.0047 | 0.0921 | 0.1047 | -0.1267 | 0.327 | 32 | 0.2983 | 1.1092 | 67.1152 | 0.0007 | 0.0117 | 0.112 | -0.0189 | 0.794 | 72 |
| yolov11x_bb_detect_model | Single-class Object Detection, Feature extraction | https://huggingface.co/IBBI-bio/ibbi_yolov11_od | https://www.arxiv.org/abs/2410.17725 | 56.9M | 640x640 | ( , 768) | 46 781 | 100 | N/A | N/A | N/A | 0.9485 | 0.9271 | 0.0741 | 1.3228 | 372.3884 | 0.0078 | 0.1096 | 0.0997 | -0.0414 | 0.756 | 32 | 0.2984 | 1.1572 | 1400.2795 | 0.0254 | 0.1364 | 0.1775 | 0.0097 | 0.916 | 72 |
| yolov10x_bb_multi_class_detect_model | Multi-class Object Detection, Feature extraction | https://huggingface.co/IBBI-bio/ibbi_yolov10_oc | https://arxiv.org/abs/2405.14458 | 29.5M | 640x640 | ( , 640) | 11 507 | 100 | 0.876 | 0.9254 | 0.2016 | 0.9593 | 0.6027 | 0.2355 | 1.1505 | 387.4236 | 0.0043 | 0.0873 | 0.1163 | -0.0891 | 0.498 | 32 | 0.2556 | 1.1246 | 46.4248 | 0.0009 | 0.0155 | 0.1122 | -0.0567 | 0.488 | 72 |
| yolov10x_bb_detect_model | Single-class Object Detection, Feature extraction | https://huggingface.co/IBBI-bio/ibbi_yolov10_od | https://arxiv.org/abs/2405.14458 | 29.5M | 640x640 | ( , 640) | 46 781 | 100 | N/A | N/A | N/A | 0.9588 | 0.9369 | 0.0436 | 1.5459 | 233.9086 | 0.0051 | 0.0826 | 0.0984 | -0.0434 | 0.742 | 32 | 0.0657 | 1.414 | 59.2355 | 0.0023 | 0.0204 | 0.1466 | 0.0324 | 0.744 | 72 |
| rtdetrx_bb_multi_class_detect_model | Multi-class Object Detection, Feature extraction | https://huggingface.co/IBBI-bio/ibbi_rtdetr_oc | https://arxiv.org/abs/2304.08069 | 76M | 640x640 | ( , 384) | 11 507 | 100 | 0.893 | 0.936 | 0.2407 | 0.9667 | 0.9167 | 0.2211 | 0.8408 | 530.0347 | 0.0065 | 0.1281 | 0.1304 | -0.139 | 0.266 | 32 | 0.3008 | 0.9058 | 115.8005 | 0.0033 | 0.0499 | 0.1163 | -0.0015 | 0.977 | 72 |
| rtdetrx_bb_detect_model | Single-class Object Detection, Feature extraction | https://huggingface.co/IBBI-bio/ibbi_rtdetr_od | https://arxiv.org/abs/2304.08069 | 76M | 640x640 | ( , 384) | 46 781 | 100 | N/A | N/A | N/A | 0.9347 | 0.9614 | 0.447 | 0.7799 | 5603.7236 | 0.0457 | 0.2687 | 0.1603 | -0.0955 | 0.486 | 32 | 0.4635 | 0.7163 | 4706.1807 | 0.0544 | 0.2874 | 0.1438 | -0.041 | 0.713 | 72 |
| grounding_dino_detect_model | Zero-shot Object Detection (Prompt: 'insect'), Feature extraction | https://huggingface.co/IDEA-Research/grounding-dino-base | https://arxiv.org/abs/2303.05499 | 341M | 640x640 | (1, 256) | 0 | N/A | N/A | N/A | N/A | 0.7238 | 0.6461 | 0.4634 | 0.7591 | 1288.2965 | 0.0146 | 0.0589 | 0.2215 | 0.0832 | 0.529 | 32 | 0.15 | 1.063 | 69.6193 | 0.0031 | 0.0396 | 0.1336 | -0.0166 | 0.87 | 72 |
| eva02_base_patch14_224_mim_in22k_features_model | Feature extraction | https://huggingface.co/timm/eva02_base_patch14_224.mim_in22k | https://arxiv.org/pdf/2303.11331 | 85.8M | 224 x 224 | (1, 768) | 0 | N/A | N/A | N/A | N/A | N/A | N/A | 0.2475 | 1.0677 | 172.0157 | 0.0005 | 0.014 | 0.0691 | -0.1673 | 0.218 | 32 | 0.3248 | 1.1139 | 137.407 | 0.0007 | 0.0074 | 0.0968 | 0.1712 | 0.092 | 72 |
| dinov3_vitl16_lvd1689m_features_model | Feature extraction | https://huggingface.co/IBBI-bio/dinov3-vitl16-pretrain-lvd1689m | https://arxiv.org/pdf/2508.10104 | 300M | 224 x 224 | ( , 384) | 0 | N/A | N/A | N/A | N/A | N/A | N/A | 0.1696 | 1.9719 | 501.7207 | 0.0036 | 0.0626 | 0.0812 | -0.1606 | 0.223 | 32 | 0.1652 | 1.5254 | 34.956 | 0.0006 | 0.0059 | 0.1064 | -0.0055 | 0.955 | 72 |
| dinov2_vitl14_lvd142m_features_model | Feature extraction | https://huggingface.co/timm/vit_large_patch14_dinov2.lvd142m | https://arxiv.org/html/2304.07193v2 | 304.5M | 336 x 336 | (1, 1024) | 0 | N/A | N/A | N/A | N/A | N/A | N/A | 0.0856 | 1.7946 | 110.4498 | 0.0004 | 0.0226 | 0.0805 | -0.0107 | 0.932 | 32 | 0.2977 | 1.3265 | 346.1827 | 0.0298 | 0.2466 | 0.2817 | 0.0548 | 0.584 | 72 |
| convformer_b36_features_model | Feature extraction | https://huggingface.co/timm/caformer_b36.sail_in22k_ft_in1k_384 | https://arxiv.org/pdf/2210.13452 | 98.8M | 384 x 384 | (1, 768) | 0 | N/A | N/A | N/A | N/A | N/A | N/A | 0.148 | 1.6396 | 84.4444 | 0.0006 | 0.0141 | 0.0627 | -0.1824 | 0.173 | 32 | 0.1927 | 1.4109 | 41.9577 | 0.0004 | 0.0042 | 0.0972 | 0.0656 | 0.525 | 72 |
For more detailed examples, please see the example notebooks located in the notebooks/ folder of the repository. Additionally the Documentation site at gcmarais.com/IBBI contains more in-depth explanations and examples.
Inference
Use inference to identify and locate bark and ambrosia beetles in an image.
# --- Create a Model ---
classifier = ibbi.create_model(model_name="species_classifier", pretrained=True)
# --- Perform Inference ---
results = classifier.predict("path/to/your/image.jpg")First, create a model using the ibbi.create_model() function. You can specify the model_name from the available models listed by ibbi.list_models(). Set pretrained=True to load the pre-trained weights.
The results from the predict() method will return a list of dictionaries containing the following keys:
labels: The predicted class label.scores: The confidence score of the prediction.boxes: The bounding box coordinates (if applicable).
NOTE: This only works for zero-shot, single-class, and multi-class detection models. Feature extraction models do not have a predict() method.
| Input Image | Single-Class Detection |
Multi-Class Species Detection |
Zero-Shot Detection |
|---|---|---|---|
 |
 |
 |
 |
Feature Extraction
All models can extract deep feature embeddings from an image. These vectors are useful for downstream tasks like clustering or similarity analysis.
# --- Create a Model ---
feature_extractor = ibbi.create_model(model_name="species_classifier", pretrained=True)
# --- Perform Inference ---
results = feature_extractor.extract_features("path/to/your/image.jpg")The results from the extract_features() method will return a NumPy array of shape (1, embedding_dimension). Different models will have different embedding dimensions, which can be found in the model summary table above.
NOTE: We recommend to create different instances of the model for inference and feature extraction to avoid any potential conflicts.
Model Evaluation
Evaluate model performance on a held-out test set using the ibbi.Evaluator() wrapper. This wrapper provides methods for computing classification, object detection, and embedding/clustering metrics to estimate the quality of feature embeddings.
⚠️ Important Note on Memory UsageThe
Evaluatormethods (.classification(),.embeddings()) currently process the entire dataset in memory. Attempting to run evaluation on the full test dataset (~2,000 images) at once may exhaust all available RAM and crash your session.To avoid this, we strongly recommend evaluating on a smaller subset of the data, as demonstrated in the code example below (using
data.select(range(10))). You can evaluate incrementally over several subsets to build a complete performance picture.
# --- Import Data ---
data = ibbi.get_dataset()
# --- Create a small subset for evaluation ---
# Use .select() to create a new Dataset object, not data[:10]
data_subset = data.select(range(10))
# --- Create a Model ---
model = ibbi.create_model(model_name="species_classifier", pretrained=True)
# --- Create an Evaluator ---
evaluator = ibbi.Evaluator(model=model, dataset=data)
# --- Classification Metrics ---
classification_results = evaluator.classification()
# --- Object Detection Metrics ---
od_results = evaluator.object_detection()
# --- Embedding & Clustering Metrics ---
embedding_results = evaluator.embeddings()You can customize the evaluation by providing your own dataset in our expected format.
Model Explainability
Understand why a model made a certain prediction. This is crucial for interpreting the model's decisions by highlighting which pixels were most influential.
# --- Create a Model ---
model = ibbi.create_model(model_name="species_classifier", pretrained=True)
# --- Create an Explainer ---
explainer = ibbi.Explainer(model=model)
# --- Explain with SHAP ---
shap_results = explainer.with_shap(explain_dataset=["path/to/your/image1.jpg", "path/to/your/image2.jpg"])
# --- Explain with LIME ---
lime_results = explainer.with_lime(image="path/to/your/image.jpg")Contributions are welcome! If you would like to improve IBBI, please see the Contribution Guide.
This project is licensed under the MIT License. See the LICENSE file for details.