GEA: Decomposing GNN embeddings with Graph Explainable Attribution
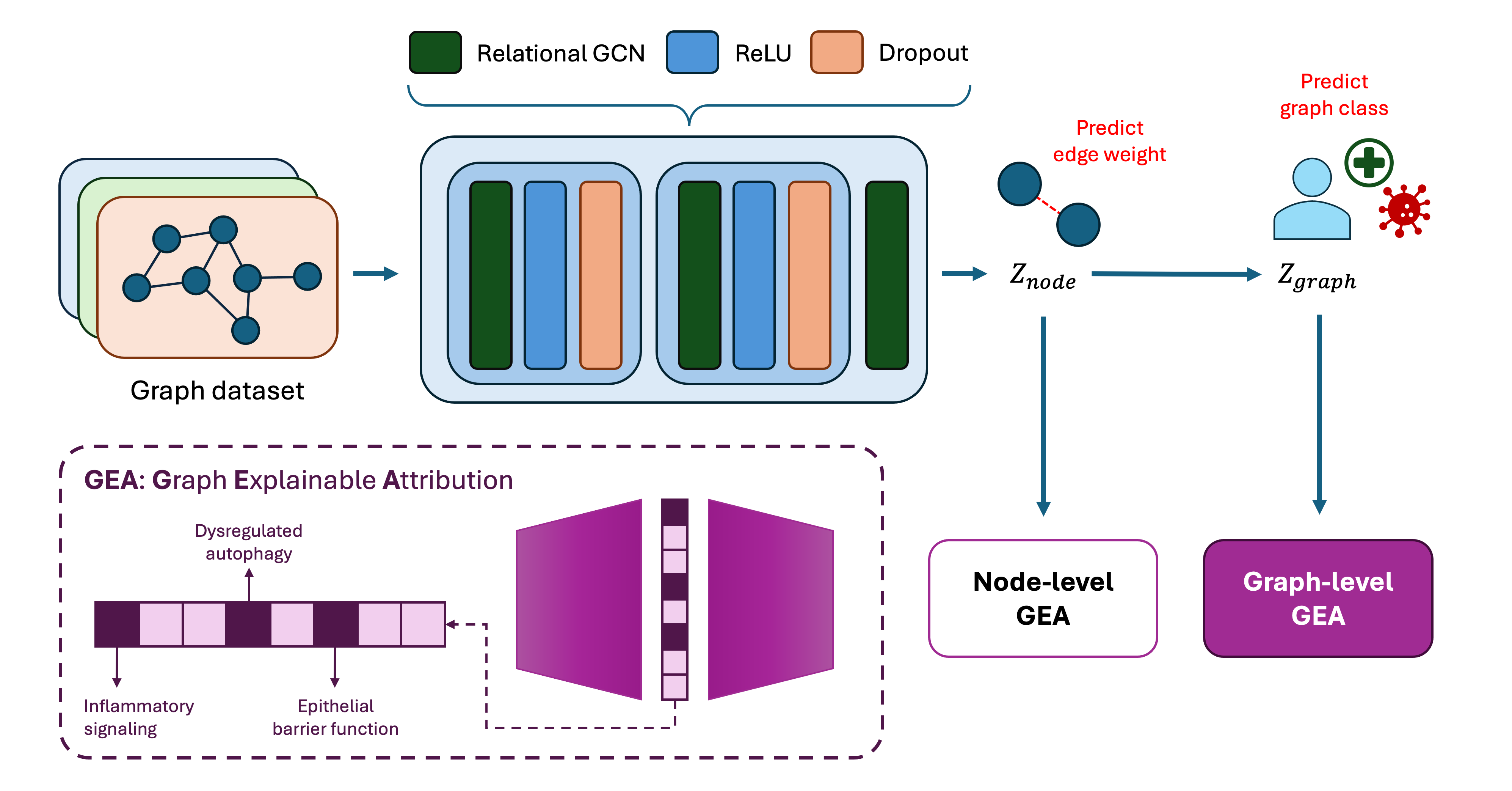
GEA is a framework for decomposing embeddings generated by a trained Graph Neural Network (GNN) model to give relevant biological insights at the node, edge and graph level. Sparse Autoencoders are employed in the framework, in which its sparse features activation can serve as actionable insights on how the internal mechanisms of the model represents information.
An overview of the workflow can be found below:
Tip
It is recommended to install GEA inside a virtual environment to manage depenendencies and avoid conflicts with existing packages. You can use the virtual environment manager of your choice, such as poetry, conda, or pipenv.
You can install the package for development by cloning this repository and running the following command:
Warning
We assume you are in the root directory of the cloned repository when running this command. Otherwise, you need to specify the path to the gea directory.
pip install -e .The code in this repository is licensed under the MIT License, allowing you to use, modify, and distribute it freely as long as you include the original copyright and license notice.