biorxiv: https://doi.org/10.1101/2023.02.18.528865
Note:
- The mouse used in the dataset is called EF65A or 65A.
- All the scripts were writen in MATLAB (i.e. 2021a), additioanl toolboxes are needed (parallel computing, image processing, curve fitting, statistics and machine learning, computer vision, bioinformatics toolbox).
- FIJI and MIJ (http://bigwww.epfl.ch/sage/soft/mij/) were used for data visualization.
- The numbers in the file names are version IDs.
- The processing and analysis pipelines are still under active development. Major changes are likely for the future axonal BARseq experiments.
- Questions and comments are welcome!
(Ongoing) deposit raw and processed data to public database
We included four variables (xyz*/mlapd*.mat) for those who want to take a quick look at the data:
- xyzDot and xyzSoma are the xyz-coordinates for rolonies (cell array) and soma (matrix) in CCFv3, with a voxel size of 25 µm.
- mlapdDot and mlapdSoma are the ML-AP-depth coordinates in the cortex for rolonies (cell array) and soma (matrix). The units for ML and AP (columns 1 and 2) are microns, and the unit for depth (column 3) is the percentage of depth within the cortical column.
Each row in these variables represents a barcoded cell, and you can locate the corresponding rolonies and soma locations using the row number. Please note that not all somas were identified due to technical issues (further details can be found in the paper), and these unidentified somas are represented as [0 0 0] in the soma variables. Additionally, only cortical rolonies were included in mlapdDot.
Example: the three signle-cell examples in Fig. 5, barcode (row) 505, 1, 431.
Necessary, contains functions for all scripts
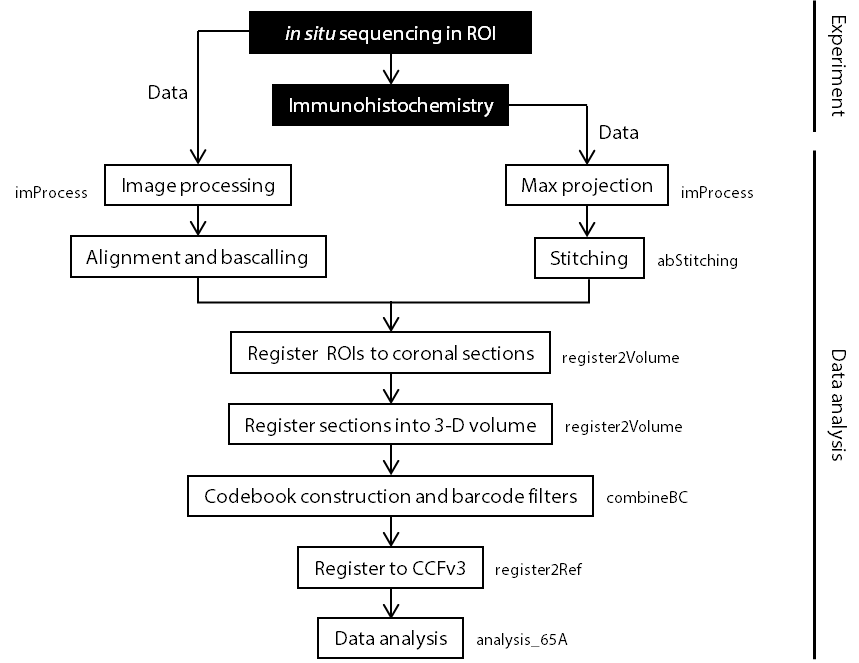
Image processing (i.e. max projection) for both sequencing and antibody data.
Processed image available, no need to rerun.
Stithing DAPI/antibody stanining whole brain images.
Processed image available, no need to rerun.
Alignment and basecalling for every batch of experiments.
Output variables available, no need to refun.
Output data need to be cleaned for the downstream analysis.
Registration experiments to a 3-D volume. Contains two parts:
- Register individual areas into whole brain coronal sections using DAPI channel.
- Register individual brain section into a 3-D volume.
Construct codebook and lookup table, exclude nonspecific barcodes.
Construct flatmap using Allen CCF (25 um/voxel).
Output: ctxAP/MP/DepthPrctile; can be directly used for 65A, no need to rerun.
Conatains two parts:
- register 3D volume to Allen CCF (Optional) Output: ref2AllenTform.mat; can be directly used for 65A, no need to rerun.
- Visualize data in registered coordinates/flatmap (contains code for Fig 1E, SupFig 4G-I)
Script for analysis of 65A data
The codes are linked to the paper but are not essential for axonal BARseq. (This part is slightly messy...)
Script for CCF-registration, by manually selecting edges of brain regions.
Output: roiCCFacc. roi selected for 65A-evaluation
Code for generating the rotating brain video/image
Code for analysis links to the Allen mouse connectivity dataset