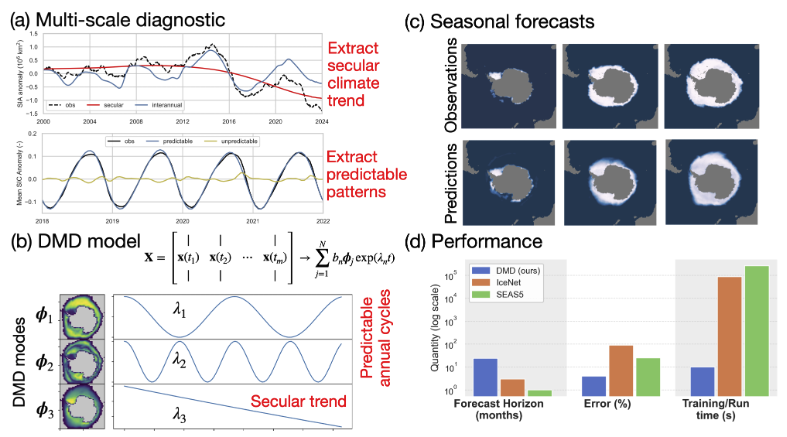
This repository details how to recreate the Figures from the manuscript "Multiscale decomposition reveals predictable interannual variability and climate trends in Antarctic sea ice loss" DOI.
- Python: 3.10+ (managed via conda — see
environment.yml) - RAM: 8 GB minimum for figure generation from precomputed results; 32 GB required for full reproduction (DMD training and precomputation)
- Disk space: ~1.5 GB for precomputed results only; ~24 GB for full raw data reproduction
- OS: Linux or macOS recommended; Windows users should use WSL
conda env create -f environment.yml
conda activate sic_dmdDownload precomputed_results.pkl from the GitHub Release page and place
it in data/. Then run any figure script:
python scripts/script_figure_6.py
python scripts/script_figure_7.py
python scripts/script_figure_8.pyRun the python script version of the mrCOSTS notebook but first set the fit flag
(line 158) to be True. Then it can be run as:
python mrCOSTS\ analysis/ice-dmd_mrcosts-figs.pyOutput of all scripts appears in figures/.
Download the files listed below and place them under data/:
| File | Size | Source |
|---|---|---|
ice_conc_sh_ease2-250_icdr-v2p0_*.nc |
5.7MB per day | OSI-SAF v2.0 (DOI) |
Antarctic_years_1989_2024i.pkl |
22 GB | Zenodo (DOI) (post-processed) |
DMD_r_5_day_34_0_thin_2_win_7.pkl |
616 MB | GitHub Release (upload in progress) |
precomputed_results.pkl |
1.3 GB | GitHub Release (upload in progress) |
coarsened-15day.all-years.v2024-09-20.nc |
36 MB | Github |
land-mask.nc |
1.5 MB | Github |
Note: The OSI SAF observation data is temporarily hosted on Dropbox via the link above. We will soon upload all data files to Zenodo and Hugging Face for permanent archival.
The precomputed file is enough to generate all predictive DMD figures.
Both DMD_r_5_day_34_0_thin_2_win_7.pkl and precomputed_results.pkl can
be regenerated from the raw observations using train_dmd.py and
precompute.py (see steps 2–3 below).
The coarsened-15day.all-years.v2024-09-20.nc and land-mask files are for the
mrCOSTS analysis. The original data were coarsened to 150 km x 150 km grids and 15 day
averages since the high-frequency, high spatial resolution components of the system
were not targetted. The land mask simply masks the location of Antarctica.
This loads the raw observations (~22 GB), builds a window-averaged training set, and trains a 100-member bootstrap DMD ensemble via BOPDMD. Requires at least 32 GB of RAM.
python scripts/train_dmd.pyOutput: data/DMD_r_5_day_34_0_thin_2_win_7.pkl (~616 MB)
This loads the raw observations and the DMD model from the previous step,
evaluates the ensemble on training and prediction windows, computes
climatology statistics, and writes data/precomputed_results.pkl.
Requires at least 32 GB of RAM.
python scripts/precompute.pyEach script loads the precomputed results and runs in a few seconds:
python scripts/script_figure_6.py # Total ice coverage prediction
python scripts/script_figure_7.py # Forecast MAE
python scripts/script_figure_8.py # Combined spatial maps + probe time series
python scripts/script_figure_s1.py # Supplementary: 2022 reconstruction
python scripts/script_figure_s2.py # Supplementary: climatology at probesThe mrCOSTS analysis is located in notebooks\ice-dmd_mrcosts-figs.ipynb and can be
run using the same environment as the predictive DMD model after setting up jupyter. See
the data notes above. You will need to make sure the conda environment is visible to
jupyter lab.
In order to reproduce the mrCOSTS figures you will need to fit mrCOSTS by setting the
fit flag to True.
The script can also be run as
python mrCOSTS\ analysis/ice-dmd_mrcosts-figs.py| Script | Paper figure | Description |
|---|---|---|
ice-dmd_mrcosts-figs.ipynb |
Figure 1 | Overview (mrCOSTS components) |
ice-dmd_mrcosts-figs.ipynb |
Figure 2 | Interannual variability (mrCOSTS) |
ice-dmd_mrcosts-figs.ipynb |
Figure 3 | Background mode spatial patterns |
ice-dmd_mrcosts-figs.ipynb |
Figure 4 | mrCOSTS band amplitudes |
ice-dmd_mrcosts-figs.ipynb |
Figure 5 | mrCOSTS point diagnosis |
script_figure_6.py |
Figure 6 | Total ice coverage: DMD vs climatology |
script_figure_7.py |
Figure 7 | Mean absolute error of DMD forecast |
script_figure_8.py |
Figure 8 | Observed vs predicted SIC maps + probe time series |
script_figure_s1.py |
Suppl. S1 | DMD reconstruction quality (2022) |
script_figure_s2.py |
Suppl. S2 | Climatology vs 2023 at probe locations |
src/sic_dmd/ Python package with shared utilities
config.py All parameters, paths, and constants
data_wrangle.py Data loading, thinning, leap-year removal
dmd_routines.py DMD training, evaluation, bootstrap
plotting.py Shared plotting helpers (colormaps, maps)
scripts/ One script per figure + DMD training + precompute
data/ Input data (not tracked in git except those used to fit mrCOSTS)
figures/ Generated figures (not tracked in git)
mrCOSTS analysis/ Notebook and script for performing the mrCOSTS analysis
If you use this code or data in your work, please cite:
@misc{yatsyshin2026multiscaledecompositionrevealspredictable,
title={Multiscale Decomposition Reveals Predictable Interannual Variability and Climate Trends in Antarctic Sea Ice Loss},
author={Peter Yatsyshin and Karl Lapo and Oliver Strickson and Louisa van Zeeland and J. Scott Hosking and J. Nathan Kutz},
year={2026},
eprint={2604.26594},
archivePrefix={arXiv},
primaryClass={physics.ao-ph},
url={https://arxiv.org/abs/2604.26594},
}